KAT3B/p300 Antibody - #AF5360
| Product: | KAT3B/p300 Antibody |
| Catalog: | AF5360 |
| Description: | Rabbit polyclonal antibody to KAT3B/p300 |
| Application: | WB IHC IF/ICC |
| Cited expt.: | WB |
| Reactivity: | Human, Mouse, Rat |
| Prediction: | Bovine, Horse, Sheep, Rabbit, Chicken, Xenopus |
| Mol.Wt.: | 300 kDa; 264kD(Calculated). |
| Uniprot: | Q09472 |
| RRID: | AB_2837845 |
Related Downloads
Protocols
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# AF5360, RRID:AB_2837845.
Fold/Unfold
E1A associated protein p300; E1A binding protein p300; E1A-associated protein p300; EP300; EP300: E1A binding protein p300; EP300_HUMAN; Histone acetyltransferase p300; KAT3B; p300 HAT; RSTS2;
Immunogens
A synthesized peptide derived from human KAT3B/p300, corresponding to a region within N-terminal amino acids.
- Q09472 EP300_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MAENVVEPGPPSAKRPKLSSPALSASASDGTDFGSLFDLEHDLPDELINSTELGLTNGGDINQLQTSLGMVQDAASKHKQLSELLRSGSSPNLNMGVGGPGQVMASQAQQSSPGLGLINSMVKSPMTQAGLTSPNMGMGTSGPNQGPTQSTGMMNSPVNQPAMGMNTGMNAGMNPGMLAAGNGQGIMPNQVMNGSIGAGRGRQNMQYPNPGMGSAGNLLTEPLQQGSPQMGGQTGLRGPQPLKMGMMNNPNPYGSPYTQNPGQQIGASGLGLQIQTKTVLSNNLSPFAMDKKAVPGGGMPNMGQQPAPQVQQPGLVTPVAQGMGSGAHTADPEKRKLIQQQLVLLLHAHKCQRREQANGEVRQCNLPHCRTMKNVLNHMTHCQSGKSCQVAHCASSRQIISHWKNCTRHDCPVCLPLKNAGDKRNQQPILTGAPVGLGNPSSLGVGQQSAPNLSTVSQIDPSSIERAYAALGLPYQVNQMPTQPQVQAKNQQNQQPGQSPQGMRPMSNMSASPMGVNGGVGVQTPSLLSDSMLHSAINSQNPMMSENASVPSLGPMPTAAQPSTTGIRKQWHEDITQDLRNHLVHKLVQAIFPTPDPAALKDRRMENLVAYARKVEGDMYESANNRAEYYHLLAEKIYKIQKELEEKRRTRLQKQNMLPNAAGMVPVSMNPGPNMGQPQPGMTSNGPLPDPSMIRGSVPNQMMPRITPQSGLNQFGQMSMAQPPIVPRQTPPLQHHGQLAQPGALNPPMGYGPRMQQPSNQGQFLPQTQFPSQGMNVTNIPLAPSSGQAPVSQAQMSSSSCPVNSPIMPPGSQGSHIHCPQLPQPALHQNSPSPVPSRTPTPHHTPPSIGAQQPPATTIPAPVPTPPAMPPGPQSQALHPPPRQTPTPPTTQLPQQVQPSLPAAPSADQPQQQPRSQQSTAASVPTPTAPLLPPQPATPLSQPAVSIEGQVSNPPSTSSTEVNSQAIAEKQPSQEVKMEAKMEVDQPEPADTQPEDISESKVEDCKMESTETEERSTELKTEIKEEEDQPSTSATQSSPAPGQSKKKIFKPEELRQALMPTLEALYRQDPESLPFRQPVDPQLLGIPDYFDIVKSPMDLSTIKRKLDTGQYQEPWQYVDDIWLMFNNAWLYNRKTSRVYKYCSKLSEVFEQEIDPVMQSLGYCCGRKLEFSPQTLCCYGKQLCTIPRDATYYSYQNRYHFCEKCFNEIQGESVSLGDDPSQPQTTINKEQFSKRKNDTLDPELFVECTECGRKMHQICVLHHEIIWPAGFVCDGCLKKSARTRKENKFSAKRLPSTRLGTFLENRVNDFLRRQNHPESGEVTVRVVHASDKTVEVKPGMKARFVDSGEMAESFPYRTKALFAFEEIDGVDLCFFGMHVQEYGSDCPPPNQRRVYISYLDSVHFFRPKCLRTAVYHEILIGYLEYVKKLGYTTGHIWACPPSEGDDYIFHCHPPDQKIPKPKRLQEWYKKMLDKAVSERIVHDYKDIFKQATEDRLTSAKELPYFEGDFWPNVLEESIKELEQEEEERKREENTSNESTDVTKGDSKNAKKKNNKKTSKNKSSLSRGNKKKPGMPNVSNDLSQKLYATMEKHKEVFFVIRLIAGPAANSLPPIVDPDPLIPCDLMDGRDAFLTLARDKHLEFSSLRRAQWSTMCMLVELHTQSQDRFVYTCNECKHHVETRWHCTVCEDYDLCITCYNTKNHDHKMEKLGLGLDDESNNQQAAATQSPGDSRRLSIQRCIQSLVHACQCRNANCSLPSCQKMKRVVQHTKGCKRKTNGGCPICKQLIALCCYHAKHCQENKCPVPFCLNIKQKLRQQQLQHRLQQAQMLRRRMASMQRTGVVGQQQGLPSPTPATPTTPTGQQPTTPQTPQPTSQPQPTPPNSMPPYLPRTQAAGPVSQGKAAGQVTPPTPPQTAQPPLPGPPPAAVEMAMQIQRAAETQRQMAHVQIFQRPIQHQMPPMTPMAPMGMNPPPMTRGPSGHLEPGMGPTGMQQQPPWSQGGLPQPQQLQSGMPRPAMMSVAQHGQPLNMAPQPGLGQVGISPLKPGTVSQQALQNLLRTLRSPSSPLQQQQVLSILHANPQLLAAFIKQRAAKYANSNPQPIPGQPGMPQGQPGLQPPTMPGQQGVHSNPAMQNMNPMQAGVQRAGLPQQQPQQQLQPPMGGMSPQAQQMNMNHNTMPSQFRDILRRQQMMQQQQQQGAGPGIGPGMANHNQFQQPQGVGYPPQQQQRMQHHMQQMQQGNMGQIGQLPQALGAEAGASLQAYQQRLLQQQMGSPVQPNPMSPQQHMLPNQAQSPHLQGQQIPNSLSNQVRSPQPVPSPRPQSQPPHSSPSPRMQPQPSPHHVSPQTSSPHPGLVAAQANPMEQGHFASPDQNSMLSQLASNPGMANLHGASATDLGLSTDNSDLNSNLSQSTLDIH
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
Functions as histone acetyltransferase and regulates transcription via chromatin remodeling. Acetylates all four core histones in nucleosomes. Histone acetylation gives an epigenetic tag for transcriptional activation. Mediates cAMP-gene regulation by binding specifically to phosphorylated CREB protein. Mediates acetylation of histone H3 at 'Lys-122' (H3K122ac), a modification that localizes at the surface of the histone octamer and stimulates transcription, possibly by promoting nucleosome instability. Mediates acetylation of histone H3 at 'Lys-27' (H3K27ac). Also functions as acetyltransferase for non-histone targets, such as ALX1, HDAC1, PRMT1 or SIRT2. Acetylates 'Lys-131' of ALX1 and acts as its coactivator. Acetylates SIRT2 and is proposed to indirectly increase the transcriptional activity of TP53 through acetylation and subsequent attenuation of SIRT2 deacetylase function. Acetylates HDAC1 leading to its inactivation and modulation of transcription. Acts as a TFAP2A-mediated transcriptional coactivator in presence of CITED2. Plays a role as a coactivator of NEUROD1-dependent transcription of the secretin and p21 genes and controls terminal differentiation of cells in the intestinal epithelium. Promotes cardiac myocyte enlargement. Can also mediate transcriptional repression. Acetylates FOXO1 and enhances its transcriptional activity. Acetylates BCL6 wich disrupts its ability to recruit histone deacetylases and hinders its transcriptional repressor activity. Participates in CLOCK or NPAS2-regulated rhythmic gene transcription; exhibits a circadian association with CLOCK or NPAS2, correlating with increase in PER1/2 mRNA and histone H3 acetylation on the PER1/2 promoter. Acetylates MTA1 at 'Lys-626' which is essential for its transcriptional coactivator activity. Acetylates XBP1 isoform 2; acetylation increases protein stability of XBP1 isoform 2 and enhances its transcriptional activity. Acetylates PCNA; acetylation promotes removal of chromatin-bound PCNA and its degradation during nucleotide excision repair (NER). Acetylates MEF2D. Acetylates and stabilizes ZBTB7B protein by antagonizing ubiquitin conjugation and degragation, this mechanism may be involved in CD4/CD8 lineage differentiation. Acetylates GABPB1, impairing GABPB1 heterotetramerization and activity (By similarity). In addition to protein acetyltransferase, can use different acyl-CoA substrates, such as (2E)-butenoyl-CoA (crotonyl-CoA), butanoyl-CoA (butyryl-CoA), 2-hydroxyisobutanoyl-CoA (2-hydroxyisobutyryl-CoA) or propanoyl-CoA (propionyl-CoA), and is able to mediate protein crotonylation, butyrylation, 2-hydroxyisobutyrylation or propionylation, respectively. Acts as a histone crotonyltransferase; crotonylation marks active promoters and enhancers and confers resistance to transcriptional repressors. Histone crotonyltransferase activity is dependent on the concentration of (2E)-butenoyl-CoA (crotonyl-CoA) substrate and such activity is weak when (2E)-butenoyl-CoA (crotonyl-CoA) concentration is low. Also acts as a histone butyryltransferase; butyrylation marks active promoters. Acts as a protein-lysine 2-hydroxyisobutyryltransferase; regulates glycolysis by mediating 2-hydroxyisobutyrylation of glycolytic enzymes. Functions as a transcriptional coactivator for SMAD4 in the TGF-beta signaling pathway. Acetylates PCK1 and promotes PCK1 anaplerotic activity. Acetylates RXRA and RXRG.
(Microbial infection) In case of HIV-1 infection, it is recruited by the viral protein Tat. Regulates Tat's transactivating activity and may help inducing chromatin remodeling of proviral genes. Binds to and may be involved in the transforming capacity of the adenovirus E1A protein.
Acetylated on Lys at up to 17 positions by intermolecular autocatalysis. Deacetylated in the transcriptional repression domain (CRD1) by SIRT1, preferentially at Lys-1020. Deacetylated by SIRT2, preferentially at Lys-418, Lys-423, Lys-1542, Lys-1546, Lys-1549, Lys-1699, Lys-1704 and Lys-1707.
Citrullinated at Arg-2142 by PADI4, which impairs methylation by CARM1 and promotes interaction with NCOA2/GRIP1.
Methylated at Arg-580 and Arg-604 in the KIX domain by CARM1, which blocks association with CREB, inhibits CREB signaling and activates apoptotic response. Also methylated at Arg-2142 by CARM1, which impairs interaction with NCOA2/GRIP1.
Sumoylated; sumoylation in the transcriptional repression domain (CRD1) mediates transcriptional repression. Desumoylated by SENP3 through the removal of SUMO2 and SUMO3.
Probable target of ubiquitination by FBXO3, leading to rapid proteasome-dependent degradation.
Phosphorylated by HIPK2 in a RUNX1-dependent manner. This phosphorylation that activates EP300 happens when RUNX1 is associated with DNA and CBFB. Phosphorylated by ROCK2 and this enhances its activity. Phosphorylation at Ser-89 by AMPK reduces interaction with nuclear receptors, such as PPARG.
Cytoplasm. Nucleus. Chromosome.
Note: Localizes to active chromatin: Colocalizes with histone H3 acetylated and/or crotonylated at 'Lys-18' (H3K18ac and H3K18cr, respectively) (PubMed:25818647). In the presence of ALX1 relocalizes from the cytoplasm to the nucleus. Colocalizes with ROCK2 in the nucleus (PubMed:12929931).
The CRD1 domain (cell cycle regulatory domain 1) mediates transcriptional repression of a subset of p300 responsive genes; it can be de-repressed by CDKN1A/p21WAF1 at least at some promoters. It conatins sumoylation and acetylation sites and the same lysine residues may be targeted for the respective modifications. It is proposed that deacetylation by SIRT1 allows sumoylation leading to suppressed activity.
Research Fields
· Cellular Processes > Cell growth and death > Cell cycle. (View pathway)
· Cellular Processes > Cellular community - eukaryotes > Adherens junction. (View pathway)
· Environmental Information Processing > Signal transduction > cAMP signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > HIF-1 signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > FoxO signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Wnt signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Notch signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > TGF-beta signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Jak-STAT signaling pathway. (View pathway)
· Human Diseases > Neurodegenerative diseases > Huntington's disease.
· Human Diseases > Infectious diseases: Bacterial > Tuberculosis.
· Human Diseases > Infectious diseases: Viral > Hepatitis B.
· Human Diseases > Infectious diseases: Viral > Influenza A.
· Human Diseases > Infectious diseases: Viral > Human papillomavirus infection.
· Human Diseases > Infectious diseases: Viral > HTLV-I infection.
· Human Diseases > Infectious diseases: Viral > Herpes simplex infection.
· Human Diseases > Infectious diseases: Viral > Epstein-Barr virus infection.
· Human Diseases > Cancers: Overview > Pathways in cancer. (View pathway)
· Human Diseases > Cancers: Overview > Viral carcinogenesis.
· Human Diseases > Cancers: Overview > MicroRNAs in cancer.
· Human Diseases > Cancers: Specific types > Renal cell carcinoma. (View pathway)
· Human Diseases > Cancers: Specific types > Prostate cancer. (View pathway)
· Organismal Systems > Nervous system > Long-term potentiation.
· Organismal Systems > Endocrine system > Melanogenesis.
· Organismal Systems > Endocrine system > Thyroid hormone signaling pathway. (View pathway)
· Organismal Systems > Endocrine system > Glucagon signaling pathway.
References
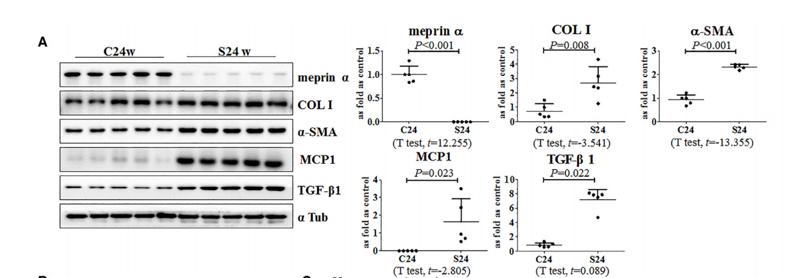
Application: WB Species: human Sample: PLC-PRF-5 cells
Application: WB Species: human Sample: MSI2 and EP300 in SW620 and LOVO cells
Application: WB Species: human Sample: myeloma cell
Application: WB Species: Rat Sample:
Application: WB Species: human Sample: ADLI cell
Application: WB Species: Mouse Sample: liver
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.