CREB Antibody - #AF6188
| Product: | CREB Antibody |
| Catalog: | AF6188 |
| Description: | Rabbit polyclonal antibody to CREB |
| Application: | WB IHC IF/ICC |
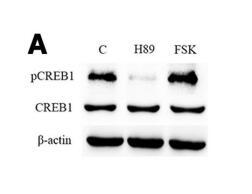
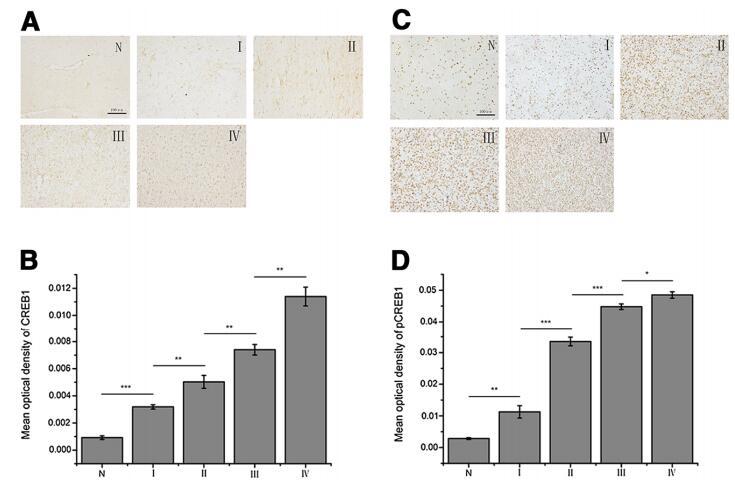
| Cited expt.: | WB, IHC |
| Reactivity: | Human, Mouse, Rat |
| Prediction: | Pig, Bovine, Horse, Sheep, Rabbit, Dog, Chicken, Xenopus |
| Mol.Wt.: | 43kDa; 37kD(Calculated). |
| Uniprot: | P16220 |
| RRID: | AB_2835071 |
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# AF6188, RRID:AB_2835071.
Fold/Unfold
Active transcription factor CREB; cAMP response element binding protein 1; cAMP response element binding protein; cAMP responsive element binding protein 1; cAMP-responsive element-binding protein 1; CREB; CREB-1; CREB1; CREB1_HUMAN; Cyclic AMP-responsive element-binding protein 1; MGC9284; OTTHUMP00000163864; OTTHUMP00000163865; OTTHUMP00000206660; OTTHUMP00000206662; OTTHUMP00000206667; Transactivator protein;
Immunogens
A synthesized peptide derived from human CREB, corresponding to a region within the internal amino acids.
- P16220 CREB1_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MTMESGAENQQSGDAAVTEAENQQMTVQAQPQIATLAQVSMPAAHATSSAPTVTLVQLPNGQTVQVHGVIQAAQPSVIQSPQVQTVQSSCKDLKRLFSGTQISTIAESEDSQESVDSVTDSQKRREILSRRPSYRKILNDLSSDAPGVPRIEEEKSEEETSAPAITTVTVPTPIYQTSSGQYIAITQGGAIQLANNGTDGVQGLQTLTMTNAAATQPGTTILQYAQTTDGQQILVPSNQVVVQAASGDVQTYQIRTAPTSTIAPGVVMASSPALPTQPAEEAARKREVRLMKNREAARECRRKKKEYVKCLENRVAVLENQNKTLIEELKALKDLYCHKSD
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
Phosphorylation-dependent transcription factor that stimulates transcription upon binding to the DNA cAMP response element (CRE), a sequence present in many viral and cellular promoters. Transcription activation is enhanced by the TORC coactivators which act independently of Ser-133 phosphorylation. Involved in different cellular processes including the synchronization of circadian rhythmicity and the differentiation of adipose cells.
Stimulated by phosphorylation. Phosphorylation of both Ser-133 and Ser-142 in the SCN regulates the activity of CREB and participates in circadian rhythm generation. Phosphorylation of Ser-133 allows CREBBP binding. In liver, phosphorylation is induced by fasting or glucagon in a circadian fashion (By similarity). CREBL2 positively regulates phosphorylation at Ser-133 thereby stimulating CREB1 transcriptional activity (By similarity). Phosphorylated upon calcium influx by CaMK4 and CaMK2 on Ser-133. CaMK4 is much more potent than CaMK2 in activating CREB. Phosphorylated by CaMK2 on Ser-142. Phosphorylation of Ser-142 blocks CREB-mediated transcription even when Ser-133 is phosphorylated. Phosphorylated by CaMK1 (By similarity). Phosphorylation of Ser-271 by HIPK2 in response to genotoxic stress promotes CREB1 activity, facilitating the recruitment of the coactivator CBP. Phosphorylated at Ser-133 by RPS6KA3, RPS6KA4 and RPS6KA5 in response to mitogenic or stress stimuli. Phosphorylated by TSSK4 on Ser-133.
Sumoylated with SUMO1. Sumoylation on Lys-304, but not on Lys-285, is required for nuclear localization of this protein. Sumoylation is enhanced under hypoxia, promoting nuclear localization and stabilization.
Nucleus.
Belongs to the bZIP family.
Research Fields
· Environmental Information Processing > Signal transduction > cGMP-PKG signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > cAMP signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > PI3K-Akt signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > AMPK signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > TNF signaling pathway. (View pathway)
· Human Diseases > Endocrine and metabolic diseases > Insulin resistance.
· Human Diseases > Neurodegenerative diseases > Huntington's disease.
· Human Diseases > Substance dependence > Cocaine addiction.
· Human Diseases > Substance dependence > Amphetamine addiction.
· Human Diseases > Substance dependence > Alcoholism.
· Human Diseases > Infectious diseases: Bacterial > Tuberculosis.
· Human Diseases > Infectious diseases: Viral > Hepatitis B.
· Human Diseases > Infectious diseases: Viral > Human papillomavirus infection.
· Human Diseases > Infectious diseases: Viral > HTLV-I infection.
· Human Diseases > Cancers: Overview > Viral carcinogenesis.
· Human Diseases > Cancers: Specific types > Prostate cancer. (View pathway)
· Organismal Systems > Aging > Longevity regulating pathway. (View pathway)
· Organismal Systems > Circulatory system > Adrenergic signaling in cardiomyocytes. (View pathway)
· Organismal Systems > Development > Osteoclast differentiation. (View pathway)
· Organismal Systems > Immune system > Antigen processing and presentation. (View pathway)
· Organismal Systems > Environmental adaptation > Circadian rhythm. (View pathway)
· Organismal Systems > Environmental adaptation > Circadian entrainment.
· Organismal Systems > Nervous system > Cholinergic synapse.
· Organismal Systems > Nervous system > Dopaminergic synapse.
· Organismal Systems > Endocrine system > Insulin secretion. (View pathway)
· Organismal Systems > Endocrine system > Estrogen signaling pathway. (View pathway)
· Organismal Systems > Endocrine system > Melanogenesis.
· Organismal Systems > Endocrine system > Thyroid hormone synthesis.
· Organismal Systems > Endocrine system > Glucagon signaling pathway.
· Organismal Systems > Endocrine system > Renin secretion.
· Organismal Systems > Endocrine system > Aldosterone synthesis and secretion.
· Organismal Systems > Endocrine system > Relaxin signaling pathway.
· Organismal Systems > Excretory system > Vasopressin-regulated water reabsorption.
References
Application: WB Species: human Sample: Gastric cancer cells
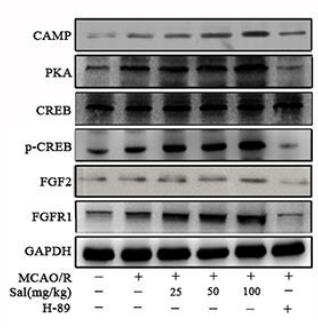
Application: WB Species: Rat Sample:
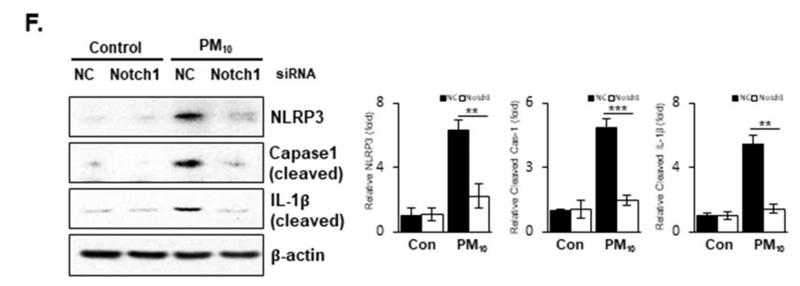
Application: WB Species: Mouse Sample:
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.