Phospho-IGF1R/Insulin Receptor (Tyr1161/Tyr1185) Antibody - #AF3125
| Product: | Phospho-IGF1R/Insulin Receptor (Tyr1161/Tyr1185) Antibody |
| Catalog: | AF3125 |
| Description: | Rabbit polyclonal antibody to Phospho-IGF1R/Insulin Receptor (Tyr1161/Tyr1185) |
| Application: | WB IHC IF/ICC IP |
| Cited expt.: | WB, IF/ICC |
| Reactivity: | Human, Mouse, Rat |
| Prediction: | Bovine, Rabbit, Dog, Chicken, Xenopus |
| Mol.Wt.: | 95kDa,155kDa; 155kD,156kD(Calculated). |
| Uniprot: | P08069 | P06213 |
| RRID: | AB_2834560 |
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# AF3125, RRID:AB_2834560.
Fold/Unfold
CD221; CD221 antigen; IGF 1 receptor; IGF 1R; IGF I receptor; IGF-I receptor; Igf1r; IGF1R_HUMAN; IGFIR; IGFIRC; IGFR; Insulin like growth factor 1 receptor; Insulin like growth factor 1 receptor precursor; Insulin-like growth factor 1 receptor beta chain; Insulin-like growth factor I receptor; JTK13; MGC142170; MGC142172; MGC18216; Soluble IGF1R variant 1; Soluble IGF1R variant 2; CD220; HHF5; HIR B; INSR; Insulin receptor; Insulin receptor subunit beta; IR;
Immunogens
A synthesized peptide derived from human IGF1R around the phosphorylation site of Tyr1161.
Found as a hybrid receptor with INSR in muscle, heart, kidney, adipose tissue, skeletal muscle, hepatoma, fibroblasts, spleen and placenta (at protein level). Expressed in a variety of tissues. Overexpressed in tumors, including melanomas, cancers of the colon, pancreas prostate and kidney.
P06213 INSR_HUMAN:Isoform Long and isoform Short are predominantly expressed in tissue targets of insulin metabolic effects: liver, adipose tissue and skeletal muscle but are also expressed in the peripheral nerve, kidney, pulmonary alveoli, pancreatic acini, placenta vascular endothelium, fibroblasts, monocytes, granulocytes, erythrocytes and skin. Isoform Short is preferentially expressed in fetal cells such as fetal fibroblasts, muscle, liver and kidney. Found as a hybrid receptor with IGF1R in muscle, heart, kidney, adipose tissue, skeletal muscle, hepatoma, fibroblasts, spleen and placenta (at protein level). Overexpressed in several tumors, including breast, colon, lung, ovary, and thyroid carcinomas.
- P08069 IGF1R_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MKSGSGGGSPTSLWGLLFLSAALSLWPTSGEICGPGIDIRNDYQQLKRLENCTVIEGYLHILLISKAEDYRSYRFPKLTVITEYLLLFRVAGLESLGDLFPNLTVIRGWKLFYNYALVIFEMTNLKDIGLYNLRNITRGAIRIEKNADLCYLSTVDWSLILDAVSNNYIVGNKPPKECGDLCPGTMEEKPMCEKTTINNEYNYRCWTTNRCQKMCPSTCGKRACTENNECCHPECLGSCSAPDNDTACVACRHYYYAGVCVPACPPNTYRFEGWRCVDRDFCANILSAESSDSEGFVIHDGECMQECPSGFIRNGSQSMYCIPCEGPCPKVCEEEKKTKTIDSVTSAQMLQGCTIFKGNLLINIRRGNNIASELENFMGLIEVVTGYVKIRHSHALVSLSFLKNLRLILGEEQLEGNYSFYVLDNQNLQQLWDWDHRNLTIKAGKMYFAFNPKLCVSEIYRMEEVTGTKGRQSKGDINTRNNGERASCESDVLHFTSTTTSKNRIIITWHRYRPPDYRDLISFTVYYKEAPFKNVTEYDGQDACGSNSWNMVDVDLPPNKDVEPGILLHGLKPWTQYAVYVKAVTLTMVENDHIRGAKSEILYIRTNASVPSIPLDVLSASNSSSQLIVKWNPPSLPNGNLSYYIVRWQRQPQDGYLYRHNYCSKDKIPIRKYADGTIDIEEVTENPKTEVCGGEKGPCCACPKTEAEKQAEKEEAEYRKVFENFLHNSIFVPRPERKRRDVMQVANTTMSSRSRNTTAADTYNITDPEELETEYPFFESRVDNKERTVISNLRPFTLYRIDIHSCNHEAEKLGCSASNFVFARTMPAEGADDIPGPVTWEPRPENSIFLKWPEPENPNGLILMYEIKYGSQVEDQRECVSRQEYRKYGGAKLNRLNPGNYTARIQATSLSGNGSWTDPVFFYVQAKTGYENFIHLIIALPVAVLLIVGGLVIMLYVFHRKRNNSRLGNGVLYASVNPEYFSAADVYVPDEWEVAREKITMSRELGQGSFGMVYEGVAKGVVKDEPETRVAIKTVNEAASMRERIEFLNEASVMKEFNCHHVVRLLGVVSQGQPTLVIMELMTRGDLKSYLRSLRPEMENNPVLAPPSLSKMIQMAGEIADGMAYLNANKFVHRDLAARNCMVAEDFTVKIGDFGMTRDIYETDYYRKGGKGLLPVRWMSPESLKDGVFTTYSDVWSFGVVLWEIATLAEQPYQGLSNEQVLRFVMEGGLLDKPDNCPDMLFELMRMCWQYNPKMRPSFLEIISSIKEEMEPGFREVSFYYSEENKLPEPEELDLEPENMESVPLDPSASSSSLPLPDRHSGHKAENGPGPGVLVLRASFDERQPYAHMNGGRKNERALPLPQSSTC
- P06213 INSR_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MATGGRRGAAAAPLLVAVAALLLGAAGHLYPGEVCPGMDIRNNLTRLHELENCSVIEGHLQILLMFKTRPEDFRDLSFPKLIMITDYLLLFRVYGLESLKDLFPNLTVIRGSRLFFNYALVIFEMVHLKELGLYNLMNITRGSVRIEKNNELCYLATIDWSRILDSVEDNYIVLNKDDNEECGDICPGTAKGKTNCPATVINGQFVERCWTHSHCQKVCPTICKSHGCTAEGLCCHSECLGNCSQPDDPTKCVACRNFYLDGRCVETCPPPYYHFQDWRCVNFSFCQDLHHKCKNSRRQGCHQYVIHNNKCIPECPSGYTMNSSNLLCTPCLGPCPKVCHLLEGEKTIDSVTSAQELRGCTVINGSLIINIRGGNNLAAELEANLGLIEEISGYLKIRRSYALVSLSFFRKLRLIRGETLEIGNYSFYALDNQNLRQLWDWSKHNLTITQGKLFFHYNPKLCLSEIHKMEEVSGTKGRQERNDIALKTNGDQASCENELLKFSYIRTSFDKILLRWEPYWPPDFRDLLGFMLFYKEAPYQNVTEFDGQDACGSNSWTVVDIDPPLRSNDPKSQNHPGWLMRGLKPWTQYAIFVKTLVTFSDERRTYGAKSDIIYVQTDATNPSVPLDPISVSNSSSQIILKWKPPSDPNGNITHYLVFWERQAEDSELFELDYCLKGLKLPSRTWSPPFESEDSQKHNQSEYEDSAGECCSCPKTDSQILKELEESSFRKTFEDYLHNVVFVPRKTSSGTGAEDPRPSRKRRSLGDVGNVTVAVPTVAAFPNTSSTSVPTSPEEHRPFEKVVNKESLVISGLRHFTGYRIELQACNQDTPEERCSVAAYVSARTMPEAKADDIVGPVTHEIFENNVVHLMWQEPKEPNGLIVLYEVSYRRYGDEELHLCVSRKHFALERGCRLRGLSPGNYSVRIRATSLAGNGSWTEPTYFYVTDYLDVPSNIAKIIIGPLIFVFLFSVVIGSIYLFLRKRQPDGPLGPLYASSNPEYLSASDVFPCSVYVPDEWEVSREKITLLRELGQGSFGMVYEGNARDIIKGEAETRVAVKTVNESASLRERIEFLNEASVMKGFTCHHVVRLLGVVSKGQPTLVVMELMAHGDLKSYLRSLRPEAENNPGRPPPTLQEMIQMAAEIADGMAYLNAKKFVHRDLAARNCMVAHDFTVKIGDFGMTRDIYETDYYRKGGKGLLPVRWMAPESLKDGVFTTSSDMWSFGVVLWEITSLAEQPYQGLSNEQVLKFVMDGGYLDQPDNCPERVTDLMRMCWQFNPKMRPTFLEIVNLLKDDLHPSFPEVSFFHSEENKAPESEELEMEFEDMENVPLDRSSHCQREEAGGRDGGSSLGFKRSYEEHIPYTHMNGGKKNGRILTLPRSNPS
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
Receptor tyrosine kinase which mediates actions of insulin-like growth factor 1 (IGF1). Binds IGF1 with high affinity and IGF2 and insulin (INS) with a lower affinity. The activated IGF1R is involved in cell growth and survival control. IGF1R is crucial for tumor transformation and survival of malignant cell. Ligand binding activates the receptor kinase, leading to receptor autophosphorylation, and tyrosines phosphorylation of multiple substrates, that function as signaling adapter proteins including, the insulin-receptor substrates (IRS1/2), Shc and 14-3-3 proteins. Phosphorylation of IRSs proteins lead to the activation of two main signaling pathways: the PI3K-AKT/PKB pathway and the Ras-MAPK pathway. The result of activating the MAPK pathway is increased cellular proliferation, whereas activating the PI3K pathway inhibits apoptosis and stimulates protein synthesis. Phosphorylated IRS1 can activate the 85 kDa regulatory subunit of PI3K (PIK3R1), leading to activation of several downstream substrates, including protein AKT/PKB. AKT phosphorylation, in turn, enhances protein synthesis through mTOR activation and triggers the antiapoptotic effects of IGFIR through phosphorylation and inactivation of BAD. In parallel to PI3K-driven signaling, recruitment of Grb2/SOS by phosphorylated IRS1 or Shc leads to recruitment of Ras and activation of the ras-MAPK pathway. In addition to these two main signaling pathways IGF1R signals also through the Janus kinase/signal transducer and activator of transcription pathway (JAK/STAT). Phosphorylation of JAK proteins can lead to phosphorylation/activation of signal transducers and activators of transcription (STAT) proteins. In particular activation of STAT3, may be essential for the transforming activity of IGF1R. The JAK/STAT pathway activates gene transcription and may be responsible for the transforming activity. JNK kinases can also be activated by the IGF1R. IGF1 exerts inhibiting activities on JNK activation via phosphorylation and inhibition of MAP3K5/ASK1, which is able to directly associate with the IGF1R.
When present in a hybrid receptor with INSR, binds IGF1.shows that hybrid receptors composed of IGF1R and INSR isoform Long are activated with a high affinity by IGF1, with low affinity by IGF2 and not significantly activated by insulin, and that hybrid receptors composed of IGF1R and INSR isoform Short are activated by IGF1, IGF2 and insulin. In contrast,shows that hybrid receptors composed of IGF1R and INSR isoform Long and hybrid receptors composed of IGF1R and INSR isoform Short have similar binding characteristics, both bind IGF1 and have a low affinity for insulin.
Autophosphorylated on tyrosine residues in response to ligand binding. Autophosphorylation occurs in trans, i.e. one subunit of the dimeric receptor phosphorylates tyrosine residues on the other subunit. Autophosphorylation occurs in a sequential manner; Tyr-1165 is predominantly phosphorylated first, followed by phosphorylation of Tyr-1161 and Tyr-1166. While every single phosphorylation increases kinase activity, all three tyrosine residues in the kinase activation loop (Tyr-1165, Tyr-1161 and Tyr-1166) have to be phosphorylated for optimal activity. Can be autophosphorylated at additional tyrosine residues (in vitro). Autophosphorylated is followed by phosphorylation of juxtamembrane tyrosines and C-terminal serines. Phosphorylation of Tyr-980 is required for IRS1- and SHC1-binding. Phosphorylation of Ser-1278 by GSK-3beta restrains kinase activity and promotes cell surface expression, it requires a priming phosphorylation at Ser-1282. Dephosphorylated by PTPN1 (By similarity).
Polyubiquitinated at Lys-1168 and Lys-1171 through both 'Lys-48' and 'Lys-29' linkages, promoting receptor endocytosis and subsequent degradation by the proteasome. Ubiquitination is facilitated by pre-existing phosphorylation.
Sumoylated with SUMO1.
Controlled by regulated intramembrane proteolysis (RIP). Undergoes metalloprotease-dependent constitutive ectodomain shedding to produce a membrane-anchored 52 kDa C-Terminal fragment which is further processed by presenilin gamma-secretase to yield an intracellular 50 kDa fragment.
Cell membrane>Single-pass type I membrane protein.
Found as a hybrid receptor with INSR in muscle, heart, kidney, adipose tissue, skeletal muscle, hepatoma, fibroblasts, spleen and placenta (at protein level). Expressed in a variety of tissues. Overexpressed in tumors, including melanomas, cancers of the colon, pancreas prostate and kidney.
Belongs to the protein kinase superfamily. Tyr protein kinase family. Insulin receptor subfamily.
Receptor tyrosine kinase which mediates the pleiotropic actions of insulin. Binding of insulin leads to phosphorylation of several intracellular substrates, including, insulin receptor substrates (IRS1, 2, 3, 4), SHC, GAB1, CBL and other signaling intermediates. Each of these phosphorylated proteins serve as docking proteins for other signaling proteins that contain Src-homology-2 domains (SH2 domain) that specifically recognize different phosphotyrosine residues, including the p85 regulatory subunit of PI3K and SHP2. Phosphorylation of IRSs proteins lead to the activation of two main signaling pathways: the PI3K-AKT/PKB pathway, which is responsible for most of the metabolic actions of insulin, and the Ras-MAPK pathway, which regulates expression of some genes and cooperates with the PI3K pathway to control cell growth and differentiation. Binding of the SH2 domains of PI3K to phosphotyrosines on IRS1 leads to the activation of PI3K and the generation of phosphatidylinositol-(3, 4, 5)-triphosphate (PIP3), a lipid second messenger, which activates several PIP3-dependent serine/threonine kinases, such as PDPK1 and subsequently AKT/PKB. The net effect of this pathway is to produce a translocation of the glucose transporter SLC2A4/GLUT4 from cytoplasmic vesicles to the cell membrane to facilitate glucose transport. Moreover, upon insulin stimulation, activated AKT/PKB is responsible for: anti-apoptotic effect of insulin by inducing phosphorylation of BAD; regulates the expression of gluconeogenic and lipogenic enzymes by controlling the activity of the winged helix or forkhead (FOX) class of transcription factors. Another pathway regulated by PI3K-AKT/PKB activation is mTORC1 signaling pathway which regulates cell growth and metabolism and integrates signals from insulin. AKT mediates insulin-stimulated protein synthesis by phosphorylating TSC2 thereby activating mTORC1 pathway. The Ras/RAF/MAP2K/MAPK pathway is mainly involved in mediating cell growth, survival and cellular differentiation of insulin. Phosphorylated IRS1 recruits GRB2/SOS complex, which triggers the activation of the Ras/RAF/MAP2K/MAPK pathway. In addition to binding insulin, the insulin receptor can bind insulin-like growth factors (IGFI and IGFII). Isoform Short has a higher affinity for IGFII binding. When present in a hybrid receptor with IGF1R, binds IGF1.shows that hybrid receptors composed of IGF1R and INSR isoform Long are activated with a high affinity by IGF1, with low affinity by IGF2 and not significantly activated by insulin, and that hybrid receptors composed of IGF1R and INSR isoform Short are activated by IGF1, IGF2 and insulin. In contrast,shows that hybrid receptors composed of IGF1R and INSR isoform Long and hybrid receptors composed of IGF1R and INSR isoform Short have similar binding characteristics, both bind IGF1 and have a low affinity for insulin. In adipocytes, inhibits lipolysis (By similarity).
After being transported from the endoplasmic reticulum to the Golgi apparatus, the single glycosylated precursor is further glycosylated and then cleaved, followed by its transport to the plasma membrane.
Autophosphorylated on tyrosine residues in response to insulin. Phosphorylation of Tyr-999 is required for binding to IRS1, SHC1 and STAT5B. Dephosphorylated by PTPRE at Tyr-999, Tyr-1185, Tyr-1189 and Tyr-1190. Dephosphorylated by PTPRF and PTPN1. Dephosphorylated by PTPN2; down-regulates insulin-induced signaling.
Cell membrane>Single-pass type I membrane protein. Late endosome. Lysosome.
Note: Binding of insulin to INSR induces internalization and lysosomal degradation of the receptor, a means for downregulating this signaling pathway after stimulation. In the presence of SORL1, internalized INSR molecules are redirected back to the cell surface, thereby preventing their lysosomal catabolism and strengthening insulin signal reception.
Isoform Long and isoform Short are predominantly expressed in tissue targets of insulin metabolic effects: liver, adipose tissue and skeletal muscle but are also expressed in the peripheral nerve, kidney, pulmonary alveoli, pancreatic acini, placenta vascular endothelium, fibroblasts, monocytes, granulocytes, erythrocytes and skin. Isoform Short is preferentially expressed in fetal cells such as fetal fibroblasts, muscle, liver and kidney. Found as a hybrid receptor with IGF1R in muscle, heart, kidney, adipose tissue, skeletal muscle, hepatoma, fibroblasts, spleen and placenta (at protein level). Overexpressed in several tumors, including breast, colon, lung, ovary, and thyroid carcinomas.
The tetrameric insulin receptor binds insulin via non-identical regions from two alpha chains, primarily via the C-terminal region of the first INSR alpha chain. Residues from the leucine-rich N-terminus of the other INSR alpha chain also contribute to this insulin binding site. A secondary insulin-binding site is formed by residues at the junction of fibronectin type-III domain 1 and 2.
Belongs to the protein kinase superfamily. Tyr protein kinase family. Insulin receptor subfamily.
Research Fields
· Cellular Processes > Cell growth and death > Oocyte meiosis. (View pathway)
· Cellular Processes > Transport and catabolism > Autophagy - animal. (View pathway)
· Cellular Processes > Transport and catabolism > Endocytosis. (View pathway)
· Cellular Processes > Cellular community - eukaryotes > Focal adhesion. (View pathway)
· Cellular Processes > Cellular community - eukaryotes > Adherens junction. (View pathway)
· Cellular Processes > Cellular community - eukaryotes > Signaling pathways regulating pluripotency of stem cells. (View pathway)
· Environmental Information Processing > Signal transduction > MAPK signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Ras signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Rap1 signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > cGMP-PKG signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > HIF-1 signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > FoxO signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > Phospholipase D signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > mTOR signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > PI3K-Akt signaling pathway. (View pathway)
· Environmental Information Processing > Signal transduction > AMPK signaling pathway. (View pathway)
· Human Diseases > Drug resistance: Antineoplastic > EGFR tyrosine kinase inhibitor resistance.
· Human Diseases > Drug resistance: Antineoplastic > Endocrine resistance.
· Human Diseases > Endocrine and metabolic diseases > Type II diabetes mellitus.
· Human Diseases > Endocrine and metabolic diseases > Insulin resistance.
· Human Diseases > Endocrine and metabolic diseases > Non-alcoholic fatty liver disease (NAFLD).
· Human Diseases > Cancers: Overview > Pathways in cancer. (View pathway)
· Human Diseases > Cancers: Overview > Transcriptional misregulation in cancer.
· Human Diseases > Cancers: Overview > Proteoglycans in cancer.
· Human Diseases > Cancers: Specific types > Glioma. (View pathway)
· Human Diseases > Cancers: Specific types > Prostate cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Melanoma. (View pathway)
· Human Diseases > Cancers: Specific types > Breast cancer. (View pathway)
· Human Diseases > Cancers: Specific types > Hepatocellular carcinoma. (View pathway)
· Organismal Systems > Aging > Longevity regulating pathway. (View pathway)
· Organismal Systems > Aging > Longevity regulating pathway - multiple species. (View pathway)
· Organismal Systems > Nervous system > Long-term depression.
· Organismal Systems > Endocrine system > Insulin signaling pathway. (View pathway)
· Organismal Systems > Endocrine system > Ovarian steroidogenesis.
· Organismal Systems > Endocrine system > Progesterone-mediated oocyte maturation.
· Organismal Systems > Endocrine system > Regulation of lipolysis in adipocytes.
· Organismal Systems > Excretory system > Aldosterone-regulated sodium reabsorption.
References
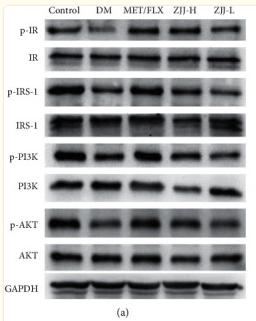
Application: WB Species: Rat Sample:
Application: WB Species: Rat Sample: hippocampus tissue
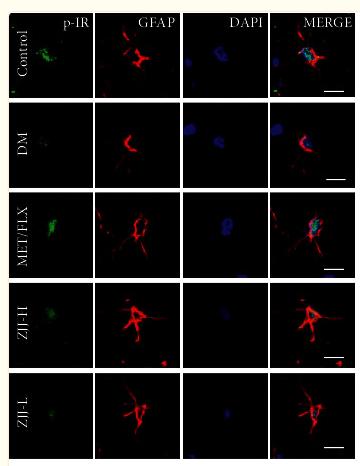
Application: IF/ICC Species: Rat Sample: hippocampus tissue
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.