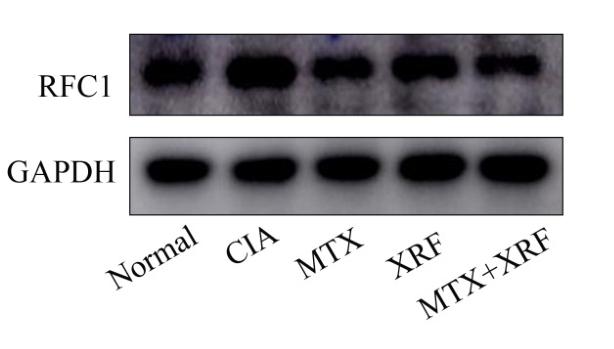
RFC1 Antibody - #DF6512
| Product: | RFC1 Antibody |
| Catalog: | DF6512 |
| Description: | Rabbit polyclonal antibody to RFC1 |
| Application: | WB IHC |
| Cited expt.: | WB |
| Reactivity: | Human, Mouse, Rat |
| Mol.Wt.: | 128kDa; 128kD(Calculated). |
| Uniprot: | P35251 |
| RRID: | AB_2838474 |
Related Downloads
Protocols
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# DF6512, RRID:AB_2838474.
Fold/Unfold
A1 140 kDa subunit; A1; A1 P145 Activator 1 large subunit; Activator 1 140 kDa subunit; Activator 1 large subunit; Activator 1 subunit 1; DNA binding Protein PO GA; DNA-binding protein PO-GA; MHC binding factor beta; MHCBFB; PO GA; RECC1; Replication factor C (activator 1) 1, 145kDa; Replication factor C 140 kDa subunit; Replication factor C; Replication factor C large subunit; Replication factor C subunit 1; Replication factor C1; RF-C 140 kDa subunit; RFC; RFC1; RFC1_HUMAN; RFC140; RFC140 Replication Factor C 140 kDa subunit;
Immunogens
A synthesized peptide derived from human RFC1, corresponding to a region within the internal amino acids.
- P35251 RFC1_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MDIRKFFGVIPSGKKLVSETVKKNEKTKSDEETLKAKKGIKEIKVNSSRKEDDFKQKQPSKKKRIIYDSDSESEETLQVKNAKKPPEKLPVSSKPGKISRQDPVTYISETDEEDDFMCKKAASKSKENGRSTNSHLGTSNMKKNEENTKTKNKPLSPIKLTPTSVLDYFGTGSVQRSNKKMVASKRKELSQNTDESGLNDEAIAKQLQLDEDAELERQLHEDEEFARTLAMLDEEPKTKKARKDTEAGETFSSVQANLSKAEKHKYPHKVKTAQVSDERKSYSPRKQSKYESSKESQQHSKSSADKIGEVSSPKASSKLAIMKRKEESSYKEIEPVASKRKENAIKLKGETKTPKKTKSSPAKKESVSPEDSEKKRTNYQAYRSYLNREGPKALGSKEIPKGAENCLEGLIFVITGVLESIERDEAKSLIERYGGKVTGNVSKKTNYLVMGRDSGQSKSDKAAALGTKIIDEDGLLNLIRTMPGKKSKYEIAVETEMKKESKLERTPQKNVQGKRKISPSKKESESKKSRPTSKRDSLAKTIKKETDVFWKSLDFKEQVAEETSGDSKARNLADDSSENKVENLLWVDKYKPTSLKTIIGQQGDQSCANKLLRWLRNWQKSSSEDKKHAAKFGKFSGKDDGSSFKAALLSGPPGVGKTTTASLVCQELGYSYVELNASDTRSKSSLKAIVAESLNNTSIKGFYSNGAASSVSTKHALIMDEVDGMAGNEDRGGIQELIGLIKHTKIPIICMCNDRNHPKIRSLVHYCFDLRFQRPRVEQIKGAMMSIAFKEGLKIPPPAMNEIILGANQDIRQVLHNLSMWCARSKALTYDQAKADSHRAKKDIKMGPFDVARKVFAAGEETAHMSLVDKSDLFFHDYSIAPLFVQENYIHVKPVAAGGDMKKHLMLLSRAADSICDGDLVDSQIRSKQNWSLLPAQAIYASVLPGELMRGYMTQFPTFPSWLGKHSSTGKHDRIVQDLALHMSLRTYSSKRTVNMDYLSLLRDALVQPLTSQGVDGVQDVVALMDTYYLMKEDFENIMEISSWGGKPSPFSKLDPKVKAAFTRAYNKEAHLTPYSLQAIKASRHSTSPSLDSEYNEELNEDDSQSDEKDQDAIETDAMIKKKTKSSKPSKPEKDKEPRKGKGKSSKK
Research Backgrounds
The elongation of primed DNA templates by DNA polymerase delta and epsilon requires the action of the accessory proteins PCNA and activator 1. This subunit binds to the primer-template junction. Binds the PO-B transcription element as well as other GA rich DNA sequences. Could play a role in DNA transcription regulation as well as DNA replication and/or repair. Can bind single- or double-stranded DNA.
Interacts with C-terminus of PCNA. 5' phosphate residue is required for binding of the N-terminal DNA-binding domain to duplex DNA, suggesting a role in recognition of non-primer template DNA structures during replication and/or repair.
Nucleus.
Wide tissue distribution. Undetectable in placental tissue.
Belongs to the activator 1 large subunit family.
Research Fields
· Genetic Information Processing > Replication and repair > DNA replication.
· Genetic Information Processing > Replication and repair > Nucleotide excision repair.
· Genetic Information Processing > Replication and repair > Mismatch repair.
References
Application: WB Species: rat Sample: intestine
Application: WB Species: rat Sample: intestine
Application: WB Species: Human Sample: HEK 293T cells
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.