HMGB1 Antibody - #AF7020
| Product: | HMGB1 Antibody |
| Catalog: | AF7020 |
| Description: | Rabbit polyclonal antibody to HMGB1 |
| Application: | WB IHC IF/ICC |
| Cited expt.: | WB, IHC, IF/ICC |
| Reactivity: | Human, Mouse, Rat |
| Prediction: | Pig, Bovine, Horse, Rabbit, Dog |
| Mol.Wt.: | 20~25kD; 25kD(Calculated). |
| Uniprot: | P09429 |
| RRID: | AB_2835324 |
Product Info
*The optimal dilutions should be determined by the end user. For optimal experimental results, antibody reuse is not recommended.
*Tips:
WB: For western blot detection of denatured protein samples. IHC: For immunohistochemical detection of paraffin sections (IHC-p) or frozen sections (IHC-f) of tissue samples. IF/ICC: For immunofluorescence detection of cell samples. ELISA(peptide): For ELISA detection of antigenic peptide.
Cite Format: Affinity Biosciences Cat# AF7020, RRID:AB_2835324.
Fold/Unfold
Amphoterin; Chromosomal protein, nonhistone, HMG1; DKFZp686A04236; High mobility group 1; High mobility group box 1; High mobility group protein 1; High mobility group protein B1; high-mobility group (nonhistone chromosomal) protein 1; HMG-1; HMG1; HMG3; HMGB 1; HMGB1; HMGB1_HUMAN; NONHISTONE CHROMOSOMAL PROTEIN HMG1; SBP 1; Sulfoglucuronyl carbohydrate binding protein;
Immunogens
A synthesized peptide derived from human HMGB1, corresponding to a region within the internal amino acids.
- P09429 HMGB1_HUMAN:
- Protein BLAST With
- NCBI/
- ExPASy/
- Uniprot
MGKGDPKKPRGKMSSYAFFVQTCREEHKKKHPDASVNFSEFSKKCSERWKTMSAKEKGKFEDMAKADKARYEREMKTYIPPKGETKKKFKDPNAPKRPPSAFFLFCSEYRPKIKGEHPGLSIGDVAKKLGEMWNNTAADDKQPYEKKAAKLKEKYEKDIAAYRAKGKPDAAKKGVVKAEKSKKKKEEEEDEEDEEDEEEEEDEEDEDEEEDDDDE
Predictions
Score>80(red) has high confidence and is suggested to be used for WB detection. *The prediction model is mainly based on the alignment of immunogen sequences, the results are for reference only, not as the basis of quality assurance.
High(score>80) Medium(80>score>50) Low(score<50) No confidence
Research Backgrounds
Multifunctional redox sensitive protein with various roles in different cellular compartments. In the nucleus is one of the major chromatin-associated non-histone proteins and acts as a DNA chaperone involved in replication, transcription, chromatin remodeling, V(D)J recombination, DNA repair and genome stability. Proposed to be an universal biosensor for nucleic acids. Promotes host inflammatory response to sterile and infectious signals and is involved in the coordination and integration of innate and adaptive immune responses. In the cytoplasm functions as sensor and/or chaperone for immunogenic nucleic acids implicating the activation of TLR9-mediated immune responses, and mediates autophagy. Acts as danger associated molecular pattern (DAMP) molecule that amplifies immune responses during tissue injury. Released to the extracellular environment can bind DNA, nucleosomes, IL-1 beta, CXCL12, AGER isoform 2/sRAGE, lipopolysaccharide (LPS) and lipoteichoic acid (LTA), and activates cells through engagement of multiple surface receptors. In the extracellular compartment fully reduced HMGB1 (released by necrosis) acts as a chemokine, disulfide HMGB1 (actively secreted) as a cytokine, and sulfonyl HMGB1 (released from apoptotic cells) promotes immunological tolerance. Has proangiogdenic activity (By similarity). May be involved in platelet activation (By similarity). Binds to phosphatidylserine and phosphatidylethanolamide (By similarity). Bound to RAGE mediates signaling for neuronal outgrowth (By similarity). May play a role in accumulation of expanded polyglutamine (polyQ) proteins such as huntingtin (HTT) or TBP.
Nuclear functions are attributed to fully reduced HGMB1. Associates with chromatin and binds DNA with a preference to non-canonical DNA structures such as single-stranded DNA, DNA-containing cruciforms or bent structures, supercoiled DNA and ZDNA. Can bent DNA and enhance DNA flexibility by looping thus providing a mechanism to promote activities on various gene promoters by enhancing transcription factor binding and/or bringing distant regulatory sequences into close proximity. May have an enhancing role in nucleotide excision repair (NER) (By similarity). However, effects in NER using in vitro systems have been reported conflictingly. May be involved in mismatch repair (MMR) and base excision repair (BER) pathways. May be involved in double strand break repair such as non-homologous end joining (NHEJ) (By similarity). Involved in V(D)J recombination by acting as a cofactor of the RAG complex: acts by stimulating cleavage and RAG protein binding at the 23 bp spacer of conserved recombination signal sequences (RSS) (By similarity). In vitro can displace histone H1 from highly bent DNA (By similarity). Can restructure the canonical nucleosome leading to relaxation of structural constraints for transcription factor-binding (By similarity). Enhances binding of sterol regulatory element-binding proteins (SREBPs) such as SREBF1 to their cognate DNA sequences and increases their transcriptional activities (By similarity). Facilitates binding of TP53 to DNA. Proposed to be involved in mitochondrial quality control and autophagy in a transcription-dependent fashion implicating HSPB1; however, this function has been questioned (By similarity). Can modulate the activity of the telomerase complex and may be involved in telomere maintenance (By similarity).
In the cytoplasm proposed to dissociate the BECN1:BCL2 complex via competitive interaction with BECN1 leading to autophagy activation. Involved in oxidative stress-mediated autophagy. Can protect BECN1 and ATG5 from calpain-mediated cleavage and thus proposed to control their proautophagic and proapoptotic functions and to regulate the extent and severity of inflammation-associated cellular injury (By similarity). In myeloid cells has a protective role against endotoxemia and bacterial infection by promoting autophagy (By similarity). Involved in endosomal translocation and activation of TLR9 in response to CpG-DNA in macrophages (By similarity).
In the extracellular compartment (following either active secretion or passive release) involved in regulation of the inflammatory response. Fully reduced HGMB1 (which subsequently gets oxidized after release) in association with CXCL12 mediates the recruitment of inflammatory cells during the initial phase of tissue injury; the CXCL12:HMGB1 complex triggers CXCR4 homodimerization. Induces the migration of monocyte-derived immature dendritic cells and seems to regulate adhesive and migratory functions of neutrophils implicating AGER/RAGE and ITGAM (By similarity). Can bind to various types of DNA and RNA including microbial unmethylated CpG-DNA to enhance the innate immune response to nucleic acids. Proposed to act in promiscuous DNA/RNA sensing which cooperates with subsequent discriminative sensing by specific pattern recognition receptors (By similarity). Promotes extracellular DNA-induced AIM2 inflammasome activation implicating AGER/RAGE. Disulfide HMGB1 binds to transmembrane receptors, such as AGER/RAGE, TLR2, TLR4 and probably TREM1, thus activating their signal transduction pathways. Mediates the release of cytokines/chemokines such as TNF, IL-1, IL-6, IL-8, CCL2, CCL3, CCL4 and CXCL10. Promotes secretion of interferon-gamma by macrophage-stimulated natural killer (NK) cells in concert with other cytokines like IL-2 or IL-12. TLR4 is proposed to be the primary receptor promoting macrophage activation and signaling through TLR4 seems to implicate LY96/MD-2. In bacterial LPS- or LTA-mediated inflammatory responses binds to the endotoxins and transfers them to CD14 for signaling to the respective TLR4:LY96 and TLR2 complexes. Contributes to tumor proliferation by association with ACER/RAGE (By similarity). Can bind to IL1-beta and signals through the IL1R1:IL1RAP receptor complex. Binding to class A CpG activates cytokine production in plasmacytoid dendritic cells implicating TLR9, MYD88 and AGER/RAGE and can activate autoreactive B cells. Via HMGB1-containing chromatin immune complexes may also promote B cell responses to endogenous TLR9 ligands through a B-cell receptor (BCR)-dependent and ACER/RAGE-independent mechanism (By similarity). Inhibits phagocytosis of apoptotic cells by macrophages; the function is dependent on poly-ADP-ribosylation and involves binding to phosphatidylserine on the cell surface of apoptotic cells (By similarity). In adaptive immunity may be involved in enhancing immunity through activation of effector T cells and suppression of regulatory T (TReg) cells. In contrast, without implicating effector or regulatory T-cells, required for tumor infiltration and activation of T-cells expressing the lymphotoxin LTA:LTB heterotrimer thus promoting tumor malignant progression (By similarity). Also reported to limit proliferation of T-cells (By similarity). Released HMGB1:nucleosome complexes formed during apoptosis can signal through TLR2 to induce cytokine production. Involved in induction of immunological tolerance by apoptotic cells; its pro-inflammatory activities when released by apoptotic cells are neutralized by reactive oxygen species (ROS)-dependent oxidation specifically on Cys-106. During macrophage activation by activated lymphocyte-derived self apoptotic DNA (ALD-DNA) promotes recruitment of ALD-DNA to endosomes (By similarity).
Phosphorylated at serine residues. Phosphorylation in both NLS regions is required for cytoplasmic translocation followed by secretion.
Acetylated on multiple sites upon stimulation with LPS. Acetylation on lysine residues in the nuclear localization signals (NLS 1 and NLS 2) leads to cytoplasmic localization and subsequent secretion (By similarity). Acetylation on Lys-3 results in preferential binding to DNA ends and impairs DNA bending activity (By similarity).
Reduction/oxidation of cysteine residues Cys-23, Cys-45 and Cys-106 and a possible intramolecular disulfide bond involving Cys-23 and Cys-45 give rise to different redox forms with specific functional activities in various cellular compartments: 1- fully reduced HMGB1 (HMGB1C23hC45hC106h), 2- disulfide HMGB1 (HMGB1C23-C45C106h) and 3- sulfonyl HMGB1 (HMGB1C23soC45soC106so).
Poly-ADP-ribosylated by PARP1 when secreted following stimulation with LPS (By similarity).
In vitro cleavage by CASP1 is liberating a HMG box 1-containing peptide which may mediate immunogenic activity; the peptide antagonizes apoptosis-induced immune tolerance. Can be proteolytically cleaved by a thrombin:thrombomodulin complex; reduces binding to heparin and proinflammatory activities (By similarity).
Nucleus. Chromosome. Cytoplasm. Secreted. Cell membrane>Peripheral membrane protein>Extracellular side. Endosome. Endoplasmic reticulum-Golgi intermediate compartment.
Note: In basal state predominantly nuclear. Shuttles between the cytoplasm and the nucleus (PubMed:12231511, PubMed:17114460). Translocates from the nucleus to the cytoplasm upon autophagy stimulation (PubMed:20819940). Release from macrophages in the extracellular milieu requires the activation of NLRC4 or NLRP3 inflammasomes (By similarity). Passively released to the extracellular milieu from necrotic cells by diffusion, involving the fully reduced HGMB1 which subsequently gets oxidized (PubMed:19811284). Also released from apoptotic cells (PubMed:16855214, PubMed:18631454). Active secretion from a variety of immune and non-immune cells such as macrophages, monocytes, neutrophils, dendritic cells and natural killer cells in response to various stimuli such as LPS and cytokines involves a nonconventional secretory process via secretory lysosomes (PubMed:12231511, PubMed:14532127, PubMed:15944249). Secreted by plasma cells in response to LPS (By similarity). Found on the surface of activated platelets (PubMed:11154118). An increased chromatin association is observed when associated with the adenovirus protein pVII (PubMed:27362237).
Ubiquituous. Expressed in platelets.
HMG box 2 mediates proinflammatory cytokine-stimulating activity and binding to TLR4 (PubMed:12765338, PubMed:20547845). However, not involved in mediating immunogenic activity in the context of apoptosis-induced immune tolerance (PubMed:24474694).
The acidic C-terminal domain forms a flexible structure which can reversibly interact intramolecularily with the HMG boxes and modulate binding to DNA and other proteins (PubMed:23063560).
Belongs to the HMGB family.
Research Fields
· Cellular Processes > Transport and catabolism > Autophagy - animal. (View pathway)
· Cellular Processes > Cell growth and death > Necroptosis. (View pathway)
· Genetic Information Processing > Replication and repair > Base excision repair.
References
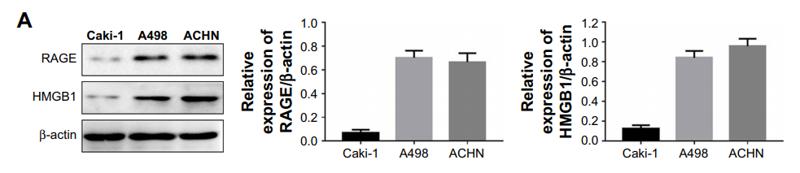
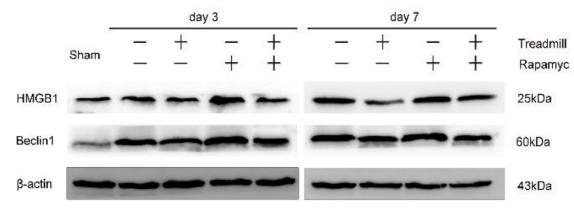
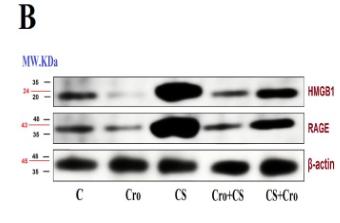
Application: IF/ICC Species: Mouse Sample: 4T1 cells
Application: IF/ICC Species: Mouse Sample: 4T1 cells
Restrictive clause
Affinity Biosciences tests all products strictly. Citations are provided as a resource for additional applications that have not been validated by Affinity Biosciences. Please choose the appropriate format for each application and consult Materials and Methods sections for additional details about the use of any product in these publications.
For Research Use Only.
Not for use in diagnostic or therapeutic procedures. Not for resale. Not for distribution without written consent. Affinity Biosciences will not be held responsible for patent infringement or other violations that may occur with the use of our products. Affinity Biosciences, Affinity Biosciences Logo and all other trademarks are the property of Affinity Biosciences LTD.











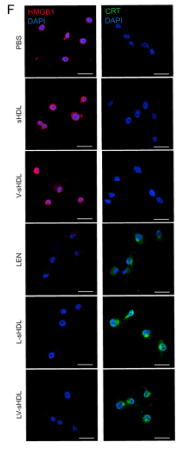


![Fig. 4. ICD and DC maturation in vitro. (A) Cell viability of CT26 cells treated with DM1, ED-sHDL, or CD-sHDL of different concentrations. (B to D) Examination of mechanisms of cell death, including apoptosis (B), necrosis (C), and ferroptosis (D). (E to G) Extracellular release of HMGB1 (E) and ATP (F) from and exposure of CRT (G) on CT26 cells after a 12-hour incubation with different reagents (1 μM in DM1). (H) Tumor-free survival of mice that had been vaccinated using differently pretreated CT26 cells (PBS, ED-sHDL, and CD-sHDL; heating, with 1 μM in DM1) after their second exposure to CT26 cancer cells (n = 5). The maturation of DC was analyzed by flow cytometry (I), and the key cytokines of DC maturation including IFN-γ, IL-12p40, and TNF-α (J) were quantified by ELISA kits after a 24-hour incubation with CT26 cells that had been pretreated with different reagents [PBS, sHDL, ED-sHDL, CD-sHDL, and lipopolysaccharide (LPS), 1 μM in DM1]. Data were presented as mean ± SD (n = 3), and statistical significance was calculated using a two-sided one-way ANOVA test. HMGB1 Antibody - Fig.](http://img.affbiotech.cn/uploads/202409/71df341c92126a7888101263b212e5c1.png)


